I am an Associate Professor at Universidade da Coruña as part of the Computer Architecture Group (GAC). Formerly I worked as a postdoc researcher in the Exelixis lab at H-ITS (Heidelberg) and I also colaborated with the Phylogenomics group (UVIGO).
The main topics of my research are Bioinformatics (molecular evolution, genomics and proteomics) and Parallel Computing for both desktop computers and cluster architectures.
Don't hesitate on asking me if you have any question about my research, or you need some advice about how not to choose the colors for your own website. I'll be glad of helping you!
BTW, my Erdös number is 4
Things I do: Things you might be interested in: (v) my punk-rock band (Morgen)
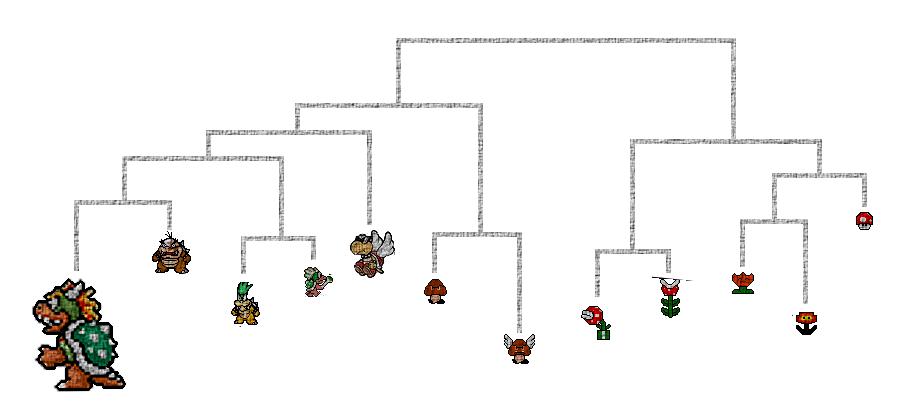
(vi) the amazing Mario Bros phylogeny (see below)
(v) my punk-rock band (Morgen)
(vi) the amazing Mario Bros phylogeny (see below)
Teaching
- Estructura de Computadores (Q3/GEI)
- Concurrencia y Paralelismo (Q4/GEI)
- Computación Paralela Avanzada (Q2/MHPC)
Research
Check my Google Scholar or ISI WoS profiles.
- D Darriba, T Flouri, A Stamatakis: The state of software for evolutionary biology. Molecular biology and evolution 35(5): 1037-1046 (2018)
- D Darriba, M Weiss, A Stamatakis: Prediction of missing sequences and branch lengths in phylogenomic data. Bioinformatics 32(9): 1331-1337 (2016)
- T Flouri, F Izquierdo-Carrasco, D Darriba, A J Aberer, L-T Nguyen, B Q Minh, A von Haeseler, A Stamatakis: The Phylogenetic Likelihood Library. Systematic Biology (2014)
- Santorum JM, Darriba D,Taboada GL, Posada D: jmodeltest.org: selection of nucleotide substitution models on the cloud. Bioinformatics 30(9): 1310-1311 (2014)
- Darriba D, Aberer A.J, Flouri T, Heath T.A, Izquierdo-Carrasco F, Stamatakis A: Boosting the performance of Bayesian divergence time estimation with the Phylogenetic Likelihood Library. 12th IEEE International Workshop on High Performance Computational Biology (HICOMB 2013). Boston, Massachusetts (USA)
- Darriba D, Taboada GL, Doallo R, Posada D: jModelTest 2: more models, new heuristics and parallel computing. Nature Methods 9:772, 2012
- Darriba D, Taboada GL, Doallo R, Posada D: ProtTest 3: fast selection of best-fit models of protein evolution. Bioinformatics 27(8): 1164-1165 (2011)
Software
Phylogenetic Likelihood Library
libPLL: The Phylogenetic Likelihood Library is a highly optimized and parallelized library for rapid prototyping and development of likelihood based phylogenetic inference codes. libPLL is the core of RAxML-NG, ModelTest-NG and EPA-NG, among others.
PLL high level modules
High level modular library for libPLL, for handling and operate with MSAs, trees, models, etc.
jModelTest 2
jModelTest is a tool to carry out statistical selection of best-fit models of nucleotide substitution. It implements five different model selection strategies: hierarchical and dynamical likelihood ratio tests (hLRT and dLRT), Akaike and Bayesian information criteria (AIC and BIC), and a decision theory method (DT). It also provides estimates of model selection uncertainty, parameter importances and model-averaged parameter estimates, including model-averaged tree topologies. jModelTest 2 includes High Performance Computing (HPC) capabilities and additional features like new strategies for tree optimization, model-averaged phylogenetic trees (both topology and branch lenght), heuristic filtering and automatic logging of user activity.
ProtTest 3
ProtTest is a bioinformatic tool for the selection of best-fit models of aminoacid replacement for the data at hand. ProtTest makes this selection by finding the model in the candidate list with the smallest Akaike Information Criterion (AIC), Bayesian Information Criterion (BIC) score or Decision Theory Criterion (DT). At the same time, ProtTest obtains model-averaged estimates of different parameters (including a model-averaged phylogenetic tree) and calculates their importance (Posada and Buckley 2004). ProtTest differs from its nucleotide analog jModeltest (Posada 2008) in that it does not include likelihood ratio tests, as not all models included in ProtTest are nested.
ForeSeqs
ForeSeqs is a tool for predicting sequences in data sets with whole missing partitions. Missing data usually lead to wrong phylogenetic inferences, since branches leading to unexistent taxa can have arbitrary lengths. Terraces in the phylogenetic tree space is another problem derived from missing taxa. Even though this problem can be avoided by linking branch lengths across partitions, the missing data can affect the inferences.
TreeChar
TreeChar is a tool for characterizing the phylogenetic tree space. Maximum-Likelihood heuristic makes assumptions on the phylogenetic tree space. However, little is known about the effect of the data characteristics on the likelihood surface, as long as it is usually not feasible to evaluate the whole tree space, which grows exponentially with the number of taxa.
Contact
- @ ddarriba [at] udc [dot] es
Computer Architecture Group
Dep. of Electronics and Systems
Facultade de Informática
Universidade de A Coruña

Phylogenomics Group
Dep. of Biochemistry, Genetics and Inmunology
Facultade de Bioloxía
Universidade de Vigo

Postal Address
Facultade de Informática
Universidad de A Coruña
Campus de Elviña s/n
15071 A Coruña
España





